[ Presentation | Mailing list | Tutorials | Get ENSnano | Pass Mac OS security check | An example ] → [2023.12.14] New version 0.5.1.1 released!
→ [2023.12.14] As Nicolas Levy left our team after briliantly defending his PhD thesis to embrace a new life at Stockly, ENSnano's git repository has moved and is now located at: https://github.com/ensnano/ensnano. Update your git and bookmarks!
ENSnano: A software for designing 3D DNA/RNA nanostructures
ENSnano takes up the concept of others DNA/RNA nanostructures design software like cadnano and scadnano, and introduces the following features:
A smooth and intuitive design interface
Fast design and handling of large structures
Custom organisation of the 2D view
3D cross-over recomandation
A new grid-based 3D organisation of helices
Realtime 3D visualisation and editing
Export of strand sequences to excel files for ordering
Import of designs from cadnano and scadnano, with 3D structure guessing and automatic grid assignement
Export to OxDNA for physical simulation, an export to MrDNA is also planed.
Cross-platform: Windows, MacOS and Linux
ENSnano is developed by Nicolas Levy and Nicolas Schabanel.
Mailing list
Join / sign off the mailing list at: https://listes.ens-lyon.fr/sympa/info/ensnano
Tutorials
The following links points to short video tutorials that will get you started with ENSnano in no time:
Getting ENSnano 0.5.1.1
Download binaries: Windows (Vulkan or DirectX), Mac OS with Intel or M1 chip
You can get binaries for Windows or MacOS on ENSnano's NEW github repository.
 Download for Windows 10 with Vulkan (preferred) |
 Download for Windows 10 for DirectX (if Vulkan version does not work only) |
 |
 |
In case mac OS blocks the app from launching, just follow the procedure to get the app approved by Mac OS:
How to pass Mac OS security check for MacOS Sonoma (MacOS 14) (Mac OS may claim that the app is "damaged" because it is not signed):
-
Open the Terminal app and type:
xattr -d com.apple.quarantine /path_to_ENSnano/ENSnano.appwhere you replace
path_to_ENSnanoby the path to the ENSnano.app (you can just typexattr -d com.apple.quarantineand then drop the ENSnano app in the terminal window, Terminal will write down the complete path for you)Note that you can then move the ENSnano app wherever you want after executing this command which must be run only once.
Compile it from source
Install Rust language. For windows, follow the link. For Linux and Mac OS, just type in a terminal:
curl --proto '=https' --tlsv1.2 -sSf https://sh.rustup.rs | shDownload the source code. For Linux and Mac OS, just type in a terminal (beware of the NEW repo direction):
git clone https://github.com/ensnano/ensnano.gitCompile and run! This will download dependencies the first time, thus it might take some time...
For Linux and Mac OS, just type in a terminal:cd ensnano cargo run --release
Changes in 0.5.1.1
- Changes in the file format, older files can still be read and will be automatically converted to the new file format when saved
Changes in 0.5.1
- Fixed a lot of typos
- Adding new curve functionality
- (Even) better DNA and RNA double-helix parameters and representation
Changes in 0.3.0
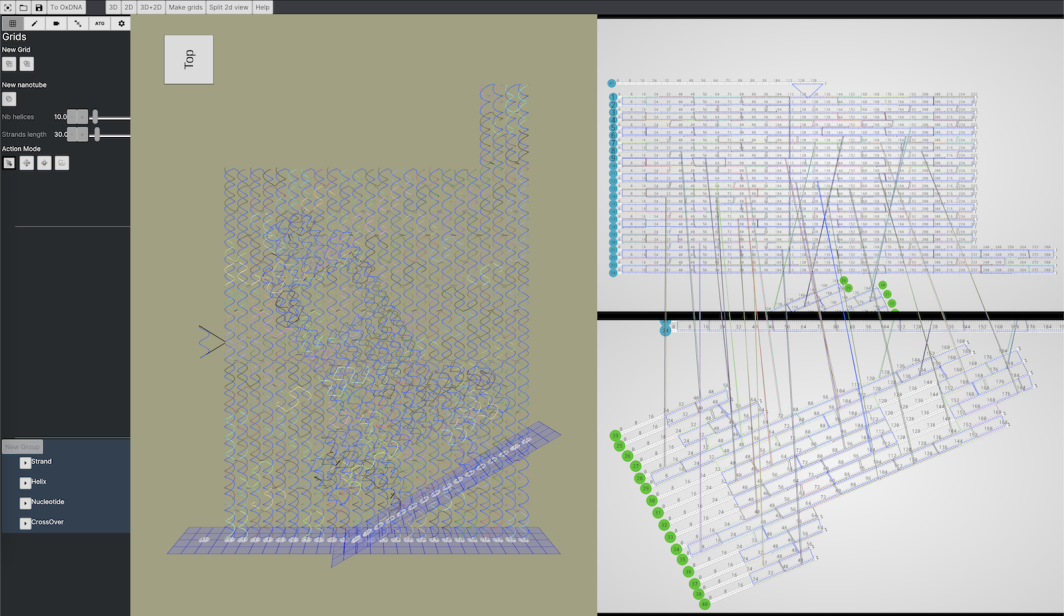
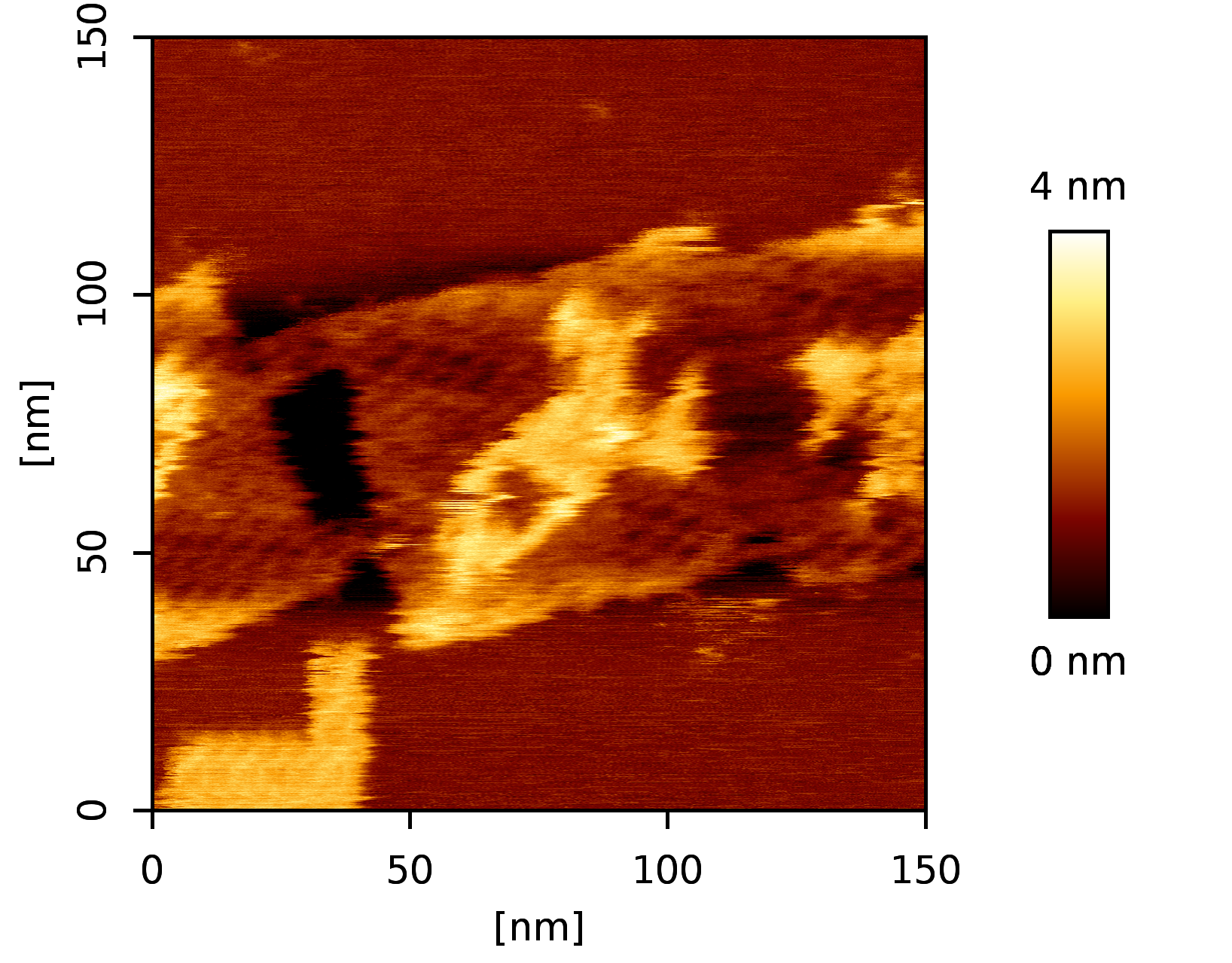
A DNA origami designed in ENSnano using 3D crossover recommendations
The following design was created in a few hours, from scratch to DNA strand order, using ENSnano.
Download the ENSnano file, staples and oxDNA export files : ENSnano-rocket_design.zip
Last update: