Condensin in the context of chromatin - P. Bernard
Our overall objective is to provide an integrative view of mitotic chromosome assembly by determining the mechanisms that make chromatin suitable for condensin action in vivo.
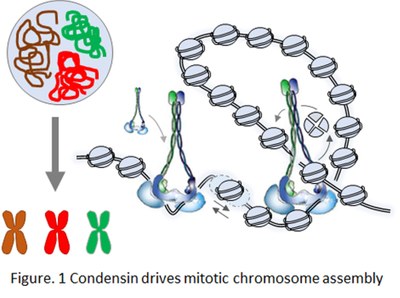 an absolute prerequisite for the accurate transmission of genomic DNA during cell division. A widely conserved, yet elusive, principle of 3D genome organisation is the looping of DNA by ring-shaped ATPase complexes of the SMC (Structural Maintenance of Chromosomes) family, such as condensin and cohesin. In vitro experiments have shown that cohesin and condensin actively extrude loops of naked DNA. Hi-C experiments confirmed that cohesin folds chromatin into loops in interphase, whilst yeast and vertebrate condensins also shape chromosomes during mitosis by forming loops and long-range intra-chromosomal contacts (Fig. 1). Yet, we still ignore: (1) the structural details of SMC-driven loop formation, and (2) how this reaction is achieved within crowded, chromatinized genomes in vivo.
an absolute prerequisite for the accurate transmission of genomic DNA during cell division. A widely conserved, yet elusive, principle of 3D genome organisation is the looping of DNA by ring-shaped ATPase complexes of the SMC (Structural Maintenance of Chromosomes) family, such as condensin and cohesin. In vitro experiments have shown that cohesin and condensin actively extrude loops of naked DNA. Hi-C experiments confirmed that cohesin folds chromatin into loops in interphase, whilst yeast and vertebrate condensins also shape chromosomes during mitosis by forming loops and long-range intra-chromosomal contacts (Fig. 1). Yet, we still ignore: (1) the structural details of SMC-driven loop formation, and (2) how this reaction is achieved within crowded, chromatinized genomes in vivo.We are specifically interested in the following questions: (1) How does condensin bind genomic DNA embedded into dense nucleosome arrays and, thus, how are chromosomal condensin binding sites defined at the chromatin level? (2) What is the impact of nucleosomes dynamics and other factors bound to chromatin, e.g. RNA polymerases and RNAs, on the DNA-looping activity of condensin and the assembly of segregation-competent mitotic chromosomes?
To tackle these fundamental questions, we searched for chromatin-associated condensin auxiliary factors through genetic and proteomic screens in fission yeast (Fig. 2). We found that specific chromatin-modifying and remodelling factors assist condensin in vivo. We proposed that fission yeast condensin binds genomic DNA in mitosis at nucleosome-poor gene promoters, possibly benefiting from the same chromatin remodelling factors as RNA Pol (Fig. 3). Yet, it remains unclear (1) if and how nucleosomes influence condensin loading in vertebrates, (2) whether or not nucleosome dynamics also impinge on the DNA looping activity of condensin, and (3) to which extent the influence of the chromosomal micro-environment on condensin vary along chromosome arms and at remarkable genomic elements such as centromeres and telomeres?
Our main objective now is to characterise the roles played by chromatin remodelling factors in the loading and functioning of condensin in vivo, by using both fission yeast and human cells as model systems.
Condensin loading during mitosis in human cultured cells
In most species, condensin massively associates with chromatin in mitosis. We proposed fission yeast suggests that nucleosomes constitute an obstacle for condensin binding to DNA that is overcome by transcription-coupled nucleosome eviction at gene promoters (Fig. 3). However, since transcription remains active during mitosis in yeasts but it is largely repressed in vertebrates, the relevance of this model to other organisms remains unclear. Human condensin I accumulates during mitosis at the promoters of genes highly expressed during the preceding G2 phase (Fig. 3). Furthermore, it is known that most active promoters remain hypersensitive to DNase I during mitosis in mammals. Thus, we speculate that persistent nucleosome depletion might shape formerly active gene promoters as loading sites for human condensin I during mitosis. We are testing this model by quantitative -MNase-seq, -ChIP-seq and degron-technology in human cultured cells. With this project, we expect to better understand how chromatin accommodates condensins in eukaryotic cells.
The impact of nucleosomes dynamics on the DNA-looping activity of condensin
Using fission yeast genetics, we collected robust evidence suggesting that nucleosome dynamics also impinge upon condensin activity in the context of chromatin. Using Hi-C, microscopy and quantitative genomics, we are further deciphering the impact of nucleosomes and their controlled turn-over on the DNA looping activity of condensin, the 3D organisation of mitotic chromosomes and their accurate transmission in anaphase.
The localisation and role of condensin at telomeres
Telomeres are the physical extremities of chromosomes. Condensin is enriched in the vicinity of telomeres in a wide-range of eukaryotic cells, and recent studies have suggested that condensin plays a role in the Telomere Dissociation Pathway (TDP) which drives sister-telomeres disjunction during anaphase (Fig. 4A). However, the mechanisms by which condensin accumulate at telomeres and take part in their TDP remains poorly understood. We identified telomere-bound factors that localise condensin at telomeres (Fig. 4B) and are studying their role with respect to condensin binding to, and activity at, telomeres in fission yeast. This work is performed in collaboration with Stéphane COULON (CRCM-Marseille) and Sylvie TOURNIER and Yannick Gachet (CBI-Toulouse).
Deciphering the link between condensin and S phase
Several lines of evidence suggest that besides its key role in chromosome assembly during mitosis, condensin might also be active on chromosomes during DNA replication and/or DNA repair, but the underlying mechanisms remain unclear. By combining genetics and proteomics approaches, performed in collaboration with the team of André Verdel in Grenoble, we identified genetic and physical links between condensin and factors involved in DNA replication/integrity during interphase (Fig. 5A). We are further studying the molecular nature of these links and their functional consequences on the organisation and transmission of chromosomes (Fig. 5B) in fission yeast.
