Replication stress and chromosome assembly and integrity-V. Vanoosthuyse
Two fundamental steps underlie the accurate transmission of the genome to daughter cells during the cell-cycle: (i) the faithful replication of DNA in S phase and (ii) the spectacular, condensin-driven reorganisation of sister-chromatids in the following mitosis, which is required for their accurate segregation to daughter cells in anaphase. Although these two essential steps are separated sometimes by several hours, they can influence each other. In particular, problems arising during DNA replication (known as DNA replication stress) can impact condensin function and the structuration and integrity of chromosomes in the following mitosis (for example, see PMID: 32966795). Such chromosome abnormalities in mitosis can contribute to aneuploidy, a hallmark of cancer. How DNA replication stress impinges on the condensin-driven assembly of chromosomes at the next mitosis remains largely unclear and is an important focus of our research.
DNA replication stress is characterised by the slowing down of replication forks and their frequent reversal and/or collapse, which increases the likelihood of deleterious DNA damage. Replication stress can be induced by the activation of oncogenes, by genome-destabilising drugs or by the depletion/mutation of specific proteins. Whatever its origin, replication stress often leads to chromosome fragility in metaphase and to chromosome segregation defects in anaphase (such as anaphase bridges or micronuclei for example). Strikingly these collective hallmarks of replication stress can be largely alleviated by the over-expression of RNase H1, a highly conserved enzyme that degrades the RNA moiety of RNA:DNA hybrids. This observation suggests that DNA replication stress disrupts the metabolism of RNA:DNA hybrids in a way that is sufficient to overwhelm the cellular machinery and to somehow trigger a wide range of chromosome abnormalities that can persist until the following mitosis. The population of stress-induced RNA:DNA hybrids with such a strong genome-destabilising impact has not yet been conclusively identified. It is also unclear whether such RNA:DNA hybrids are merely a toxic, stress-induced byproduct of fork transactions that needs to be quickly removed, or whether they are an important, albeit short-lived, intermediary in a genome-stabilising process. A major objective of our research is to identify and characterise these stress-induced RNA:DNA hybrids and to decipher how they impact chromosome integrity.
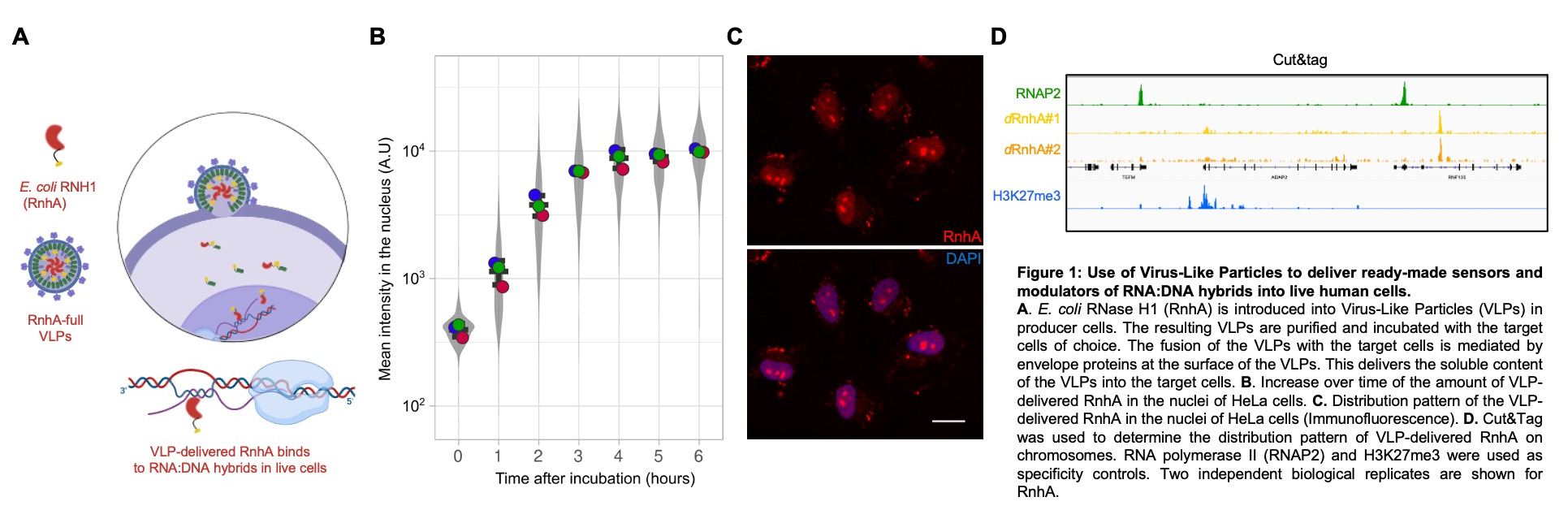
A better understanding of the genome-destabilising impact of RNA:DNA hybrids requires new tools to quickly manipulate RNA:DNA hybrids in live cells. We have developed an innovative approach to map and manipulate RNA:DNA hybrids in live human cells. We use Virus-Like Particles to deliver a panel of ready-made regulators and sensors of RNA:DNA hybrids directly into live cells. Our results show that such delivery is quick and happens under four hours at physiologically-relevant levels (Fig 1). Such a quick delivery is compatible with the manipulation and the mapping of RNA:DNA hybrids in specific phases of the cell-cycle. We aim to use these new innovative tools on synchronous populations of human cells in culture to identify and characterise the population of genome-destabilising RNA:DNA hybrids that is triggered upon replication stress.
For all enquiries, please contact vincent.vanoosthuyse [at] ens-lyon.fr
Il n'y a aucun élément dans ce dossier pour l'instant.