Publication by CIRI on Nucleic Acids Research on October 10, 2019.
Abstract
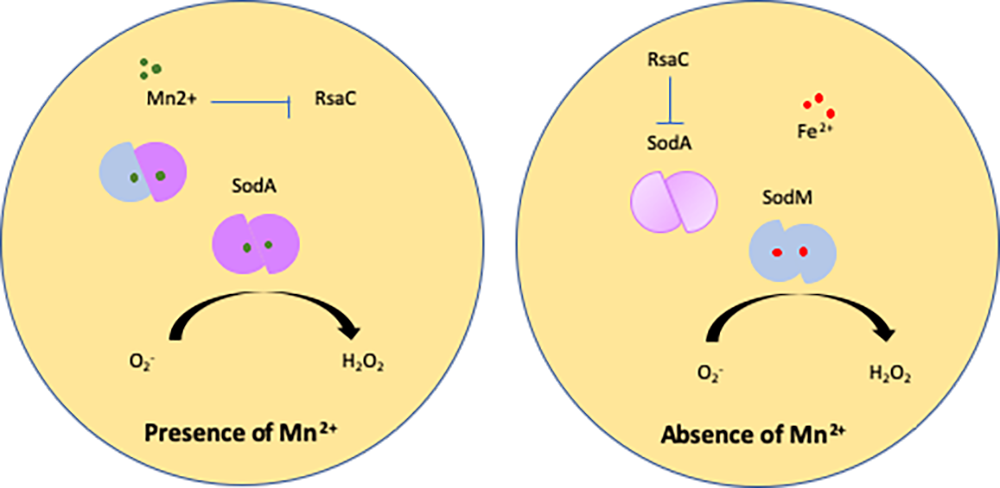 The human opportunistic pathogen Staphylococcus aureus produces numerous small regulatory RNAs (sRNAs) for which functions are still poorly understood. Here, we focused on an atypical and large sRNA called RsaC. Its length varies between different isolates due to the presence of repeated sequences at the 5' end while its 3' part is structurally independent and highly conserved. Using MS2-affinity purification coupled with RNA sequencing (MAPS) and quantitative differential proteomics, sodA mRNA was identified as a primary target of RsaC sRNA. SodA is a Mn-dependent superoxide dismutase involved in oxidative stress response. Remarkably, rsaC gene is co-transcribed with the major manganese ABC transporter MntABC and, consequently, RsaC is mainly produced in response to Mn starvation. This 3'UTR-derived sRNA is released from mntABC-RsaC precursor after cleavage by RNase III. The mature and stable form of RsaC inhibits the synthesis of the Mn-containing enzyme SodA synthesis and favors the oxidative stress response mediated by SodM, an alternative SOD enzyme using either Mn or Fe as co-factor. In addition, other putative targets of RsaC are involved in oxidative stress (ROS and NOS) and metal homeostasis (Fe and Zn). Consequently, RsaC may balance two interconnected defensive responses, i.e. oxidative stress and metal-dependent nutritional immunity.
The human opportunistic pathogen Staphylococcus aureus produces numerous small regulatory RNAs (sRNAs) for which functions are still poorly understood. Here, we focused on an atypical and large sRNA called RsaC. Its length varies between different isolates due to the presence of repeated sequences at the 5' end while its 3' part is structurally independent and highly conserved. Using MS2-affinity purification coupled with RNA sequencing (MAPS) and quantitative differential proteomics, sodA mRNA was identified as a primary target of RsaC sRNA. SodA is a Mn-dependent superoxide dismutase involved in oxidative stress response. Remarkably, rsaC gene is co-transcribed with the major manganese ABC transporter MntABC and, consequently, RsaC is mainly produced in response to Mn starvation. This 3'UTR-derived sRNA is released from mntABC-RsaC precursor after cleavage by RNase III. The mature and stable form of RsaC inhibits the synthesis of the Mn-containing enzyme SodA synthesis and favors the oxidative stress response mediated by SodM, an alternative SOD enzyme using either Mn or Fe as co-factor. In addition, other putative targets of RsaC are involved in oxidative stress (ROS and NOS) and metal homeostasis (Fe and Zn). Consequently, RsaC may balance two interconnected defensive responses, i.e. oxidative stress and metal-dependent nutritional immunity.
Source: RsaC sRNA modulates the oxidative stress response of Staphylococcus aureus during manganese starvation. Lalaouna D, Baude J, Wu Z, Tomasini A, Chicher J, Marzi S, Vandenesch F, Romby P, Caldelari I, Moreau K. Nucleic Acids Res. 10 octobre 2019 ; 47(18):9871-9887. DOI : 10.1093/nar/gkz728.






